Epigenomics of clear cell renal cell carcinoma: mechanisms and potential use in molecular pathology
Kidney cancer, now being the 7th leading cancer in men in US, contributing to the estimated 61,560 newly diagnosed and 14,080 deaths in 2015 (1). Renal cell carcinoma (RCC) is the most common form of kidney cancer of which 70–80% of cases are defined as clear cell RCC (ccRCC), which is the RCC subtype we discuss below. The incidence of RCC has been steadily rising by 2–4% each year in recent decades worldwide. Even though the incidence of RCC in Asia is lower than in US and Europe, the mortality-to-morbidity ratio is much higher in Asia than in Western countries (2). In China, there is obvious increment of the RCC incidence in recent years with the early diagnosed cases increasing.
Epigenetics is a phenomenon that involves heritable changes in gene expression that does not affect the nucleotide sequence but impacts the conformation of the DNA (3). Epigenetics crucially determine which genes are expressed by which cell type, and when (4). Epigenetic alterations are considered as a hallmark of cancer (5). At least three types of epigenetic modifications regulate chromatin: DNA methylation, histone modifications, and non-coding RNAs. The most well documented epigenetic alterations are DNA methylation and histone modifications. Unlike genetic alteration, such epigenetic changes can potentially be reversed, making enzymes involved in such processes promising therapeutic targets. RCC is characterized by numerous genetic and epigenetic alterations. Here we reviewed the important epigenetic alterations in DNA methylation, histone modifications and non-coding RNAs including microRNAs and recently identified long noncoding RNAs during RCC tumorigenesis and progression, especially their potential uses in molecular diagnosis, prognostic and possible therapeutic prediction.
DNA methylation in ccRCC
DNA methylation and tumorigenesis of ccRCC
DNA methylation includes hypermethylation of area rich in cytosine and guanine dinucleotides (CpG islands, CpGIs) within the promoter that results in tumor suppressor gene (TSG) inactivation and affects noncoding RNA expression or promoter CpGIs hypomethylation resulting in proto-oncogene activation. Recent data have shown that the non-CpGIs loci are also very important in gene regulation. Furthermore, newer high-resolution assays reveal that gene body methylation may be even more important in gene regulation than promoter methylation.
It is well known that the familial and nearly 70% of sporadic ccRCC cases are characterized by the alterations of von Hippel–Lindau (VHL) gene resulting from either somatic mutation in majority status, loss of heterozygosity and promoter hypermethylation (6). The dysfunction of VHL leads to the accumulation of hypoxia-inducible factors (HIFs) and increased effects of angiogenesis and pathogenesis of ccRCC (7,8). Although the VHL inactivation by promoter methylation occurs in ~15% of sporadic ccRCC, inactivation of the TSG in RCC most commonly results from promoter methylation, whilst intragenic mutations are rare.
By comparing DNA methylation profiles (1,505 CpGIs analyzed) between familial and sporadic VHL wild type ccRCC cases, several loci, including RASSF1, PITX2, CDH13, HS3ST2, TWIST1, TAL1, TUSC3, and DCC were found to be more frequently methylated in sporadic VHL wild-type ccRCC than in familial ccRCCs (9). This finding might suggest that DNA methylation complicated the ccRCC tumorigenesis pathway(s) and give possible explanation of low effectiveness in cancer therapy if only blocking the downstream effects of VHL inactivation.
Numerous genes have been found to be frequently methylated in ccRCC by candidate-gene approaches (50% of cases), including CDH1, APAF1, COL1A1, DKK2, DKK3, SFRP1, SFRP4, SFRP5, WIF, PCDH17 and TCF21, which are not methylated or are rarely methylated in matched normal renal tissues (<10%) (10-12). These genes are involved in different aspects of tumorigenesis, such as signal transduction, apoptosis, angiogenesis, adhesion and tumor invasion. Another recent comparative study of CpG methylation status in 38 ccRCCs and 9 matched normal kidney tissues including 27,500 CpGs at >14,000 genes showed that 55 genes were identified as methylated in ccRCC but not in normal controls. Further detailed functional study revealed eight novel ccRCC TSG candidates, including OVOL1, DLEC1, BMP4, SST, TMPRSS2, TM6SF1, SLC34A2, and COL1A2. Among them, OVOL1 epigenetic silencing was identified to be capable of increasing c-Myc expression and may lead to c-Myc pathway activation commonly seen in ccRCC formation (13).
Several functional epigenetic strategies were applied for ccRCC TSGs identification. One strategy is RCC cell lines treated with 5-aza-2-deoxycytidine were subjected to high-density gene expression microarray analysis. The re-expressed genes after demethylation were further validated in primary tumor samples. Five genes including BNC1, COL14A1, CST6, PDLIM4, and SFRP1 were revealed to show frequent (>30%) promoter region methylation associated with transcriptional silencing. Further the cell growth assay in vitro by over expression or RNAi knock down of these genes suggested their tumor suppressor activities (12). The other strategy is methylated DNA immunoprecipitation (MeDIP) in combination with high-density whole-genome expression microarray for direct analysis of genomic methylation patterns in primary ccRCC. Nine genes, including ATP5G2, PCDH8, CORO6, KLHL35, QPCT, SCUBE3, ZSCAN18, CCDC8, and FBN2 were found frequently methylated in primary ccRCC and promoter hypermethylation of these genes resulted in significant reduction of their expression level (14).
Using the HELP (HpaII tiny fragment enrichment by ligation-mediated PCR) assay, a very recent study determined the CpG methylation status of 1.3 million loci across the genome in RCC and found that the aberrant methylation was particularly enriched in kidney-specific enhancer regions. These aberrantly hypermethylated regions revealed enrichment for binding sites of AP2a, AHR, HAIRY, ARNT, and HIF1 transcription factors, contributing to the dysregulated hypoxia signaling pathways in RCC (15).
DNA methylation and prognosis of ccRCC
Genome-scale methylation analysis is a powerful tool to identify multi-gene profiles reflecting tumor behavior, supported by a comprehensive methylome analysis using single-CpG resolution infinium array. This study has revealed ccRCCs with positive CpGI methylator phenotype on the FAM150A, GRM6, ZNF540, ZFP42, ZNF154, RIMS4, PCDHAC1, KHDRBS2, ASCL2, KCNQ1, PRAC, WNT3A, TRH, FAM78A, ZNF671, SLC13A5 and NKX6-2 genes were characterized by more aggressive tumor phenotypes and poorer patient outcomes (16).
The very recent high-resolution methylome analysis of RCC demonstrated that only methylation at enhancers was highly prognostic for survival in even multivariate analysis (15).
DNA methylation and molecular diagnosis of ccRCC
Some promoter hypermethylation have been detected in serum, peripheral blood and urine, suggesting the blood and body fluid-based DNA methylation candidate marker’s potential value in early non-invasive detection of ccRCC (10,17). For example, the levels of promoter hypermethylation of RASSF1A (Ras association domain family member 1A) and VHL detected in serum of ccRCC patients (18), as well as that of KILLIN (a novel p53-regulated TSG proximal to PTEN) and long interspersed nuclear elements (LINE-1) detected in peripheral blood (19,20), were significantly higher than that in patients with benign tumors and healthy controls, respectively. Certainly the practical significance of these methylation candidate markers deserves validation in large, independent cohorts.
So far, there is no data available for DNA methylation markers in prediction of chemotherapeutic response in RCC patients, even though several differentially methylated genes were proposed as candidate markers of drug response in human cancer (21). ‘Epigenetic treatment’, with DNA methyltransferases inhibition and silenced gene activity restoration, resulting in enhanced apoptosis and decreased cancer cell growth, has become a promising therapeutic strategy for ccRCC. This novel strategy may reprogram the gene expression profiles in cancer cells thereby making ccRCC more susceptible to standard therapy (22).
Histone modifications in ccRCC
DNA, histones and nonhistone proteins are condensed into a highly complex nucleoprotein structure known as chromatin, either in a compact heterochromatin form or a more open euchromatin form, which is related to transcriptionally active genes. Histones on the N-terminus modifications include methylation, acetylation, phosphorylation, ubiquitylation and sumoylation of specific residues, of which histone acetylation and methylation are the most studied (23). Generally, acetylation causes active transcription and is associated with a more open chromatin conformation, whereas the transcriptional effects of methylation depend on the residue affected as well as the degree of methylation (mono-, di- or trimethylation). For example, methylation of H3K4, H3K26 and H3K79 is associated with active marks, whereas methylation of H3K9, H3K27 and H4K20 is associated with repressive marks (24).
Histone deacetylase (HDACs), histone acetyltransferases (HATs), histone methyltransferases (HMTs) and recently discovered histone demethylases (HDMTs) are critical enzymes in regulating various biological and cellular processes including cell proliferation, angiogenesis, hypoxia-related effects and cell cycle regulation. Increasing studies have found the deregulation of these enzymes in ccRCC. Targeting these enzymes, reversing their epigenetic modifications has the potential of reactivating TSGs or suppressing oncogenes, thus, affecting tumor growth or progression of ccRCC.
Histone modification, especially histone-modifying enzymes in ccRCC initiation and progression
HIF isoforms are shown to be correlated with histone-modifying enzymes. The interaction between HDAC4 and HIF-1α could protect HIF-α from proteasomal degradation (25). JARID1C, which encodes a HIF-targeted histone H3K4 demethylase and has truncating mutations in RCC (3%), is believed to be a TSG in VHL null tumors (26). JMJD1A, one HMT transcriptionally induced by HIF2α under hypoxic conditions, is consistent overexpressed in RCC. Several studies have reported that SETD2, which encodes a histone H3K36 methyltransferase located (3p21.31) in close proximity to VHL gene within a commonly lost (approximately 90%) 3p locus, as a novel TSG in ccRCC (27). The somatic truncating mutations in SETD2 (3–8%) were associated with VHL mutations (28), hypoxic phenotype and decline in chromatin remodeling complex protein PBRM1 in RCC tumor samples, as well as advanced stage (29). These results indicate that histone-modifying enzymes are especially important in RCC with VHL mutation or deletions.
Epithelial-to-mesenchymal transition (EMT) is believed to play an important role in ccRCC progression, as well as in many other cancers. HDAC inhibition could suppress TGF-β-induced EMT of renal proximal tubular epithelial cells (30), thereby may indicate a potential role for HDAC inhibitor in reversing EMT during RCC progression, which deserve further exploration.
HDACs are also essential in the regulation of the cell cycle and apoptosis through their effects on pRB and p53. When combined with other agents, such as rapamycin (an inhibitor of mTORC1), HDACs inhibitors have been shown to have strong anti-tumor activity in xenografts of certain RCC cell types (31).
Enhancer of zeste homolog 2 (EZH2), the core catalytic subunit of polycomb repressive complex (PRC2), acts as a HMT by adding 3 methylgroups to lysine 27 of histone 3 (H3K27). The trimethylation of H3K27 leads to chromatin condensation and mediates epigenetic silencing of a broad range of TSG involved in cancer progression. EZH2 could promote migration and invasion of RCC cells via E-cadherin repression (32). MiR-101 act as a negative regulator of EZH2 expression (33).
Histone modification and prognosis of ccRCC
The level of histone H4 acetylation (H4Ac) was inversely correlated with the pathological stage and nuclear grade of ccRCC, whereas lower H3Ac levels were associated with systemic metastatic spread and tumor progression. Furthermore, the low H3K18Ac level (lysines 18 in histone H3) was significantly correlated with RCC progression in univariate analysis and it could be used as an independent predictor of cancer progression and poor survival following surgery in localized RCC (34).
A series of reports have showed that the lower methylation levels of histone 3 in different lysine loci were of worse prognostic value. The lower levels of global H3K4 methylation (mono-, di- and trimethylation), especially H3K4me2, in RCC tumor samples were reported to be correlated with the adverse clinicopathological parameters such as more advanced pathological stage and higher Fuhrman grade as well as worse progression-free survival or cancer-specific survival in univariate analysis; suggesting they could be used as a predictor of poor prognosis of RCC (35,36). The methylation H3K9 level was also found to be of prognostic significance in RCC patients. The lower H3K9me2 level was shown to be a significant predictor of worse outcome in RCC even in multivariate analysis including nuclear grade, tumor location and the putative biomarkers (Ki67 index and p53) (34). The lower level of global methylation levels of lysine 27 of histone H3 (H3K27me1, H3K27me2 and H3K27me3) were shown to be associated with advanced pathological stage, higher Fuhrman grade and vascular invasion. In addition, lower H3K27me3 level was observed in patients with distant metastasis. Furthermore progression-free survival was shorter in patients with lower H3K27me1 and H3K27me3 levels, although only in a univariate analysis (37).
Numerous reports have showed that EZH2 overexpression was an adverse prognostic biomarker in different types of carcinomas. So far limited studies have suggested that the higher intratumoral expression of EZH2 was correlated with the advanced stages and poor survival of ccRCC patients both in primary and metastatic cases (38,39), however the conflicting results exist.
miRNA and ccRCC
MiRNAs are a group of small (around 22 nucleotides) non-coding RNAs regulating post-transcriptional gene expression through the epigenetic mechanism of RNA interference (40). The 6 to 8 nucleotides at 5' end of a miRNA known as seed region, can determine the specificity of its interaction with the target mRNA on its 3' untranslated region, resulting in the inhibition of translation and/or degradation of mRNA (41,42). The abnormal expression of miRNAs is related to cancer development, progression and metastasis. Overexpressed miRNAs in cancer may act like oncogenes by down-regulating TSG. Vice versa, down-regulated tumor suppressor-like miRNAs could result in the up-regulation of oncogenes. In general the oncogenic miRNAs are upregulated and tumor suppressor miRNAs are downregulated in malignancy. Accumulating evidences show that miRNAs play important roles in the tumorigenesis and progression of RCC (43).
miRNA and initiation of ccRCC
The role of miRNA in the VHL-HIF pathway and their relationship have been of particular interest and explored in many reports. MiR-17-5p, miR-224, miR-200 family, miR106a/b, miR-21, miR-221, miR-199a, and miR-214 have been identified to have prominent roles in ccRCC formation. They are predicted to target the VHL-HIF1 axis along with the targets of HIF/HIF transcription factors: vascular epithelial growth factor (VEGF), platelet-derived growth factor, transforming growth factor, the downstream PI3K/AKT and RAS/RAF/MEK/ERK pathways, and mTOR pathway. The same miRNA can control multiple targets along the same pathway, and multiple miRNAs can target the same molecule. This implies that even subtle alterations in these miRNAs may cause significant influences on the outcome of the regulated pathway and the tumorigenesis of ccRCC. This also indicates that miRNAs can control the hypoxia-related VHL-HIF1 axis and the common cancer-related pathways in a tight regulatory network.
MiR-17-92 cluster are oncogenic miRNAs in many cancers (44). MiR-92a, being a member of miR-17-92 cluster, is found to be overexpressed in ccRCC and correlated negatively with VHL mRNA levels (45). Meanwhile, the presence of direct interactions between miR-17-5p (also a member of miR-17-92 cluster), miR-224 with VHL and HIF1-α was validated by luciferase assay and Western blot analysis, as well as in ccRCC specimens. In addition, miR-17-5p can regulate multiple key components of hypoxia-related signaling pathways, including VEGFA and ENLN3. MiR-224 can inhibit the TGF-β pathway members SMAD4 and SMAD5 (46). MiR-210 is the most ubiquitously upregulated miRNA in multiple cancer cell lines under hypoxia. As a result of HIF1 or HIF2 over expression in ccRCC cells, miR-210 is found to be upregulated and target ISCU (iron sulphur cluster homologue), which acts as a scaffold protein for the formation of Fe-S clusters which may contribute to anaerobic respiration in tumors (47). Moreover, the accumulation of miR-210 may induce aneuploidy via E2F3 down regulation and contribute to the initiation of ccRCC (48). Besides the HIF pathway, dysfunction of VHL also works through other ways. One of major VHL functions is inhibiting beta-catenin. In VHL inactivated RCC, beta-catenin is the direct target of miR-1826 and they show an inverse correlation (49). In another report, increased expression of miR-28-5p could promote chromosome instability by inhibiting mitotic checkpoint protein Mad2 in VHL inactivated tumors (50). However, so far few studies report how miRNAs function in VHL independent ccRCCs.
miRNA and progression of ccRCC
The invasion and metastasis of cancer start with the enhanced ability of tumor cell to destruct extracellular matrix (ECM) proteins and invade ECM, which can be assisted by metastasis promoting miRNAs (51). MiR-21 is overexpressed in ccRCC and contributes to ccRCC invasion and metastasis. Studies revealed miR-21 could inhibit the tumor suppressor PDCD4. Overexpressed miR-21 results in the phosphorylation and activation of AKT and IKKβ, further activation of NF-kB dependent signal pathway transcription. Together with its blocking PDCD4 mediated the inhibitory effects; miR-21 overexpression could restore the invasion and migration of ccRCC cells. Moreover, IKKβ activates mTOR1, which further restores cell invasion and migration (52). EMT is a crucial process involved in tumor motility and invasion. Down regulation of several miRNAs has been shown to cause EMT in ccRCC (53). Tumor suppressors miR-200 family (miR-200a/b/c, miR-141, miR-429) are significantly downregulated in RCC cells, which repress the expression of E-cadherin through direct targeting their transcriptional inhibitor ZEB1 and ZEB2 (54,55). For miR-141, its decrease can activate the EphA2/p-FAK/p-AKT/MMPs signaling cascade to facilitate the invasion of RCC cells (56). The process of EMT can also be triggered by HIF dependent down-regulation of miR-30C, which leads to subsequent increase of slug and decrease of E-cadherin (57). Furthermore, tumor suppressive miR-138 is significantly reduced in ccRCC. Through its targeting of vimentin, a significantly overexpressed intermediate filament protein in ccRCC tissues, miR-138 could contribute to cell migration and invasion (58). CAV2, a member of caveolin family, function as a signal transduction regulator in the invasion and migration of ccRCC cells. Tumor suppressor miR-218 is reported to directly regulate CAV2. Downregulation of miR-218 and up-regulation of CAV2 might be frequent events in ccRCC (59).
Some well-defined miRNAs and their targets in ccRCC are summarized in Table 1. The regulatory functions of these important miRNAs in the major signaling pathways of ccRCC initiation and progression are illustrated in Figure 1.
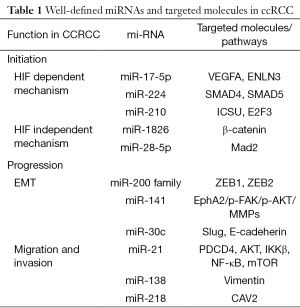

Full table
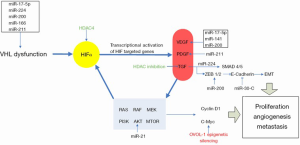
miRNA and molecular diagnosis of ccRCC
MicroRNAs are highly stable and abundant in plasma, serum and other body fluids. They can passively leak from damaged or apoptotic cells into circulation. Some miRNAs are contained within the extracellular vesicles, thus resistant to RNase and persistent in blood. Detection and characterization of circulating miRNAs can be a promising non-invasive way to detect cancer (60,61). Recent studies suggested miR-210 as a potential biomarker for early diagnosis of ccRCC. As we mentioned before, the elevation of miR-210 could be resulted from the over activation of VHL-HIF pathway. As in cancer tissues, miR-210 level in serum was found to be increased significantly in RCC patients compared to non-cancer controls (62). Another study found that the level of miR-378 was increased and miR-451 level was decreased in the serum of RCC patients. These two miRNAs combination enable the identification of RCC in serum with the sensitivity of 81% and specificity of 83% (63). A panel of 5 serum miRNAs, including three significantly increased miRNAs (miR-193a-3p, miR-362 and miR-572) and two markedly decreased miRNAs (miR-28-5p and miR-378), can differentiate stage I RCC from non-cancer controls, suggesting it may have the potential to be used clinically as an auxiliary tool for the early detection of RCC (64).
Analysis of urinary miRNAs is another promising approach for early diagnosis of urological cancer (65), although there are limited studies available for RCC. Same as in blood, the miRNA level in urine also shows stability (66). Different from the usual down-regulation of miR-15a in other cancers, miR-15a is upregulated in ccRCC biopsies and urine samples, which is not seen in other urinary and nonurinary tumors, and inflammatory conditions of the urinary tract. The finding suggests miR-15a can be a potential RCC biomarker (67).
By integrating the miRNAs, mRNAs, and pathways, some miRNAs are suggested to be the novel ccRCC biomarkers, such as miR-425, miR-136, miR-335, miR-340 and miR-320d (68). So far none of these markers are identified to be specific for RCC. For instance, miR-210 and miR-425 aberrations are also detected in carcinomas of digestive system.
miRNA and prognosis of ccRCC
Many studies have investigated the relationship between the alteration of miRNAs and prognosis of ccRCC (Table 2). The aberrant expression of miRNAs is frequently related to poorer prognosis, because tumor suppressor miRNAs are usually downregulated and pro-oncogenic miRNAs are upregulated. Up-regulation of miR-21 is a marker for poor prognosis in several cancers including RCC (69). Furthermore, the ratio of miR-21/10b was shown to be an independent prognostic marker for metastasis-free ccRCC (70). For the tumor suppressing miRNAs, studies have showed that the decreased expression of miR-497, miR-23b-27b cluster are associated with the advanced stage and shorter overall survival of RCC patients (71,72). As for miR-210, its overexpression was demonstrated as a poor prognosis marker for RCC in one study (73), while a conflict result relating to good prognosis was shown in another study (47).
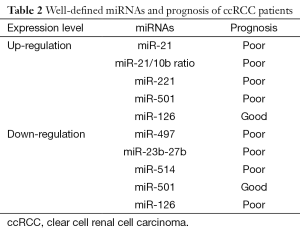
Full table
MiR-501 is differently expressed in RCC, regulating the balance of cell apoptosis and proliferation. Its high expression level predicts a worse outcome and vice versa (74). Similarly, miR-126 can also be differently expressed, however its down-regulation is related to the poor prognosis and its up-regulation is associated with significantly prolonged disease-free survival of RCC patients (75).
By comparing the miRNA expression in primary ccRCC and metastatic ccRCC, miR-122 was found to be upregulated in primary tumor and reduced in metastatic tumor. MiR-514 was shown to be downregulated in primary tumor and decreased further in metastatic tumor. These miRNA alterations are significantly related to recurrence risk and miR-514 could provide independent prognostic information for ccRCC patients (76).
Patients with higher plasma miR-221 levels had a significantly lower survival rate than those with lower expression levels, revealing miR-221 as an independent prognosis factor for ccRCC.
LncRNAs and ccRCC
Long non-coding RNAs (lncRNAs) are operationally defined as non-protein coding RNAs that are over 200 nucleotides in length, which corresponds to a convenient cutoff in biochemical fractionation and excludes all known small RNAs (77). LncRNAs are divided into two main categories, the intergenic lncRNA and the intragenic lncRNA. They are differentially expressed in tissue specific manner. Many lncRNAs undergo 5' end capping and 3' end polyadenylation similar to mRNAs. They were once considered as transcriptional noise while accumulated evidences show that lncRNAs have a regulatory role in cellular development, proliferation, differentiation by cis-regulation and trans-regulation (78). And their misregulation could cause various human diseases including cancer (79). Studies have revealed the cancer-involved functions of lncRNAs as oncogenes, tumor suppressor, or both.
LncRNAs and progression of ccRCC
lncRNA MALAT1, located on chromosome 11q13, is expressed ubiquitously in normal tissues (80). Several studies found that the expression of MALAT1 was elevated in multiple cancers such as non-small cell lung carcinoma, hepatocellular carcinoma, and pancreatic carcinoma, suggesting MALAT1 acting as an oncogene. It could enhance cell growth, tumorigenesis and metastasis by activating E2F1 transcriptional factor or ERK/MAPK pathway (81,82). MALAT1 was also upregulated significantly in ccRCC compared to the adjacent normal tissue. Further, the up-regulation of MALAT1 was related to tumor size, stage and lymph node metastasis but not histological grade and distant metastasis, which prompted MALAT1 overexpression was correlated with ccRCC progression and development (83). HOTAIR is another oncogenic lncRNA that interacts with the PRC and suppresses its target genes. HOTAIR is upregulated in several types of cancers and stimulates cancer development (84-86). The knockdown of HOTAIR could inhibit the proliferation, migration, and invasive ability of RCC cells. In addition, inhibition of this lncRNA by siRNA suppressed tumor formation in vivo (87). One recent study showed the expression and function of HOTAIR could be regulated by tumor suppressor miR-141. MiR-141 binds to HOTAIR in a sequence specific manner and induces the cleavage mediated by Ago2 complex (88). lncRNA SPRY4-IT1 was found to be elevated significantly in several types of cancers including ccRCC. Moreover its high expression was shown to be associated with advanced histological grade, tumor stage and lymph node or distant metastasis. Knocking down its expression could inhibit the proliferation, migration and invasion of ccRCC cells. All these findings indicate the oncogenic role of lncRNA SPRY4-IT1 in ccRCC (89). For the tumor suppressor lncRNAs, lncRNA GAS5 was found to be decreased significantly in ccRCC both in vitro and in vivo. Its normal expression suppressed the invasion and migration of ccRCC cell. However, GAS5 expression did not show any correlation with the clinicopathological factors (90). Another studied tumor suppressing lncRNA is CADM1-AS1, which is located at the antisense direction of a coding exon of CADM1. CADM1 is a tumor suppressor that encodes a cellular adhesion molecule and regulates cell cycle, apoptosis and differentiation (91). The study revealed that lncRNA CADM1-AS1 expression was significantly associated with the CADM1 mRNA expression and reversely correlated with the RCC stage (92).
LncRNAs and molecular classification of ccRCC
By next-generation deep RNA-sequencing of the 475 primary ccRCC samples from The Cancer Genome Atlas (TCGA), lncRNA expressions were assessed and the first lncRNA based RCC classification was proposed in a recent study. Totally 1,934 expressed lncRNA were identified. Unsupervised clustering analysis unraveled four lncRNA subclasses in ccRCC associated with distinct clinicopathological and genomic features. The cluster C2 is enriched for 9p deletion, chr8q24.22 gain and chromatin remodeler BAP1 somatic mutations. Bearing frequent 1p deletion, 17q gain, and MiTF/TFE translocations, the cluster C4 (7.8%) is related to tumor subtypes arising from the distal tubules of the nephron, which is proved to be chromophobe RCC, clear cell papillary RCC and translocation associated RCC (92).
LncRNAs and prognosis of ccRCC
In the lncRNA subtype classification of ccRCC we mentioned above, cluster C2 (23.4%) defines the most aggressive tumors, with the highest Fuhrman grade, stage and the worst overall survival (92). The oncogenic lncRNAs are usually upregulated and related to poor prognosis in RCC. MALAT1 and SPRY4-IT1 showed the potential to become the powerful independent prognostic factor for RCC (83,91). Down-regulation of tumor suppressor lncRNA CADM1-AS1 is related to worse prognosis compared to patients with higher expression level of CADM1-AS1 (92). The above limited findings are listed in Table 3.
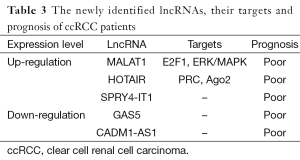
Full table
Epigenomics and target therapy for ccRCC
The recent advances in the clarification of molecular mechanisms involved in ccRCC have led to the development and approval of multiple agents targeting VEGF, such as bevacizumab plus interferon, sorafenib, sunitinib, pazopanib and axitinib, or mTOR pathways, such as temsirolimus and everolimus for treatment of advanced ccRCC (93). The epigenomic studies, especially inhibiting HDACs and reversing promoter methylation of those important ccRCC TSG, are providing promising targets for novel therapies. Several agents targeting Class I, II and IV HDACs have been tested in preclinical studies, functioning as a single agent or in combination with proteasome inhibitor bortezomib, temsirolimus, interferon or IL-2 etc. Furthermore, demethylating agents such as azacitidine and decitabine have been shown to be useful for the treatment of advanced ccRCC besides its successful utility in myelodysplastic syndrome and acute myelogenous leukemia.
In conclusion, epigenetic alterations regulating the formation and progression of RCC are still in the initiating stage of discovery. Further characterization of the genes and pathways that are epigenetically altered in ccRCC may lead to the development of novel minimally-invasive diagnostic and prognostic tools for ccRCC. In the long term, epigenetic therapies might provide an additional treatment option for advanced RCC that is unresponsive to standard management.
Acknowledgements
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin 2015;65:5-29. [PubMed]
- Patel AR, Prasad SM, Shih YC, et al. The association of the human development index with global kidney cancer incidence and mortality. J Urol 2012;187:1978-83. [PubMed]
- Bird A. DNA methylation patterns and epigenetic memory. Genes Dev 2002;16:6-21. [PubMed]
- Beyond the genome. Nature 2015;518:273. [PubMed]
- Sandoval J, Esteller M. Cancer epigenomics: beyond genomics. Curr Opin Genet Dev 2012;22:50-5. [PubMed]
- Bausch B, Jilg C, Gläsker S, et al. Renal cancer in von Hippel-Lindau disease and related syndromes. Nat Rev Nephrol 2013;9:529-38. [PubMed]
- Kim W, Kaelin WG Jr. The von Hippel-Lindau tumor suppressor protein: new insights into oxygen sensing and cancer. Curr Opin Genet Dev 2003;13:55-60. [PubMed]
- Linehan WM, Rubin JS, Bottaro DP. VHL loss of function and its impact on oncogenic signaling networks in clear cell renal cell carcinoma. Int J Biochem Cell Biol 2009;41:753-6. [PubMed]
- McRonald FE, Morris MR, Gentle D, et al. CpG methylation profiling in VHL related and VHL unrelated renal cell carcinoma. Mol Cancer 2009;8:31. [PubMed]
- Rydzanicz M, Wrzesiński T, Bluyssen HA, et al. Genomics and epigenomics of clear cell renal cell carcinoma: recent developments and potential applications. Cancer Lett 2013;341:111-26. [PubMed]
- Costa VL, Henrique R, Danielsen SA, et al. TCF21 and PCDH17 methylation: An innovative panel of biomarkers for a simultaneous detection of urological cancers. Epigenetics 2011;6:1120-30. [PubMed]
- Morris MR, Maher ER. Epigenetics of renal cell carcinoma: the path towards new diagnostics and therapeutics. Genome Med 2010;2:59. [PubMed]
- Ricketts CJ, Morris MR, Gentle D, et al. Genome-wide CpG island methylation analysis implicates novel genes in the pathogenesis of renal cell carcinoma. Epigenetics 2012;7:278-90. [PubMed]
- Morris MR, Ricketts CJ, Gentle D, et al. Genome-wide methylation analysis identifies epigenetically inactivated candidate tumour suppressor genes in renal cell carcinoma. Oncogene 2011;30:1390-401. [PubMed]
- Hu CY, Mohtat D, Yu Y, et al. Kidney cancer is characterized by aberrant methylation of tissue-specific enhancers that are prognostic for overall survival. Clin Cancer Res 2014;20:4349-60. [PubMed]
- Arai E, Chiku S, Mori T, et al. Single-CpG-resolution methylome analysis identifies clinicopathologically aggressive CpG island methylator phenotype clear cell renal cell carcinomas. Carcinogenesis 2012;33:1487-93. [PubMed]
- Baldewijns MM, van Vlodrop IJ, Schouten LJ, et al. Genetics and epigenetics of renal cell cancer. Biochim Biophys Acta 2008;1785:133-55.
- de Martino M, Klatte T, Haitel A, et al. Serum cell-free DNA in renal cell carcinoma: a diagnostic and prognostic marker. Cancer 2012;118:82-90. [PubMed]
- Bennett KL, Campbell R, Ganapathi S, et al. Germline and somatic DNA methylation and epigenetic regulation of KILLIN in renal cell carcinoma. Genes Chromosomes Cancer 2011;50:654-61. [PubMed]
- Liao LM, Brennan P, van Bemmel DM, et al. LINE-1 methylation levels in leukocyte DNA and risk of renal cell cancer. PLoS One 2011;6:e27361. [PubMed]
- Heyn H, Esteller M. DNA methylation profiling in the clinic: applications and challenges. Nat Rev Genet 2012;13:679-92. [PubMed]
- Negrotto S, Hu Z, Alcazar O, et al. Noncytotoxic differentiation treatment of renal cell cancer. Cancer Res 2011;71:1431-41. [PubMed]
- Bannister AJ, Kouzarides T. Regulation of chromatin by histone modifications. Cell Res 2011;21:381-95. [PubMed]
- Ramakrishnan S, Ellis L, Pili R. Histone modifications: implications in renal cell carcinoma. Epigenomics 2013;5:453-62. [PubMed]
- Qian DZ, Kachhap SK, Collis SJ, et al. Class II histone deacetylases are associated with VHL-independent regulation of hypoxia-inducible factor 1 alpha. Cancer Res 2006;66:8814-21. [PubMed]
- Niu X, Zhang T, Liao L, et al. The von Hippel-Lindau tumor suppressor protein regulates gene expression and tumor growth through histone demethylase JARID1C. Oncogene 2012;31:776-86. [PubMed]
- Duns G, van den Berg E, van Duivenbode I, et al. Histone methyltransferase gene SETD2 is a novel tumor suppressor gene in clear cell renal cell carcinoma. Cancer Res 2010;70:4287-91. [PubMed]
- Dalgliesh GL, Furge K, Greenman C, et al. Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 2010;463:360-3. [PubMed]
- Hakimi AA, Chen YB, Wren J, et al. Clinical and pathologic impact of select chromatin-modulating tumor suppressors in clear cell renal cell carcinoma. Eur Urol 2013;63:848-54. [PubMed]
- Yoshikawa M, Hishikawa K, Marumo T, et al. Inhibition of histone deacetylase activity suppresses epithelial-to-mesenchymal transition induced by TGF-beta1 in human renal epithelial cells. J Am Soc Nephrol 2007;18:58-65. [PubMed]
- Verheul HM, Salumbides B, Van Erp K, et al. Combination strategy targeting the hypoxia inducible factor-1 alpha with mammalian target of rapamycin and histone deacetylase inhibitors. Clin Cancer Res 2008;14:3589-97. [PubMed]
- Liu L, Xu Z, Zhong L, et al. Enhancer of zeste homolog 2 (EZH2) promotes tumour cell migration and invasion via epigenetic repression of E-cadherin in renal cell carcinoma. BJU Int 2016;117:351-62. [PubMed]
- Sakurai T, Bilim VN, Ugolkov AV, et al. The enhancer of zeste homolog 2 (EZH2), a potential therapeutic target, is regulated by miR-101 in renal cancer cells. Biochem Biophys Res Commun 2012;422:607-14. [PubMed]
- Seligson DB, Horvath S, McBrian MA, et al. Global levels of histone modifications predict prognosis in different cancers. Am J Pathol 2009;174:1619-28. [PubMed]
- Mosashvilli D, Kahl P, Mertens C, et al. Global histone acetylation levels: prognostic relevance in patients with renal cell carcinoma. Cancer Sci 2010;101:2664-9. [PubMed]
- Ellinger J, Kahl P, Mertens C, et al. Prognostic relevance of global histone H3 lysine 4 (H3K4) methylation in renal cell carcinoma. Int J Cancer 2010;127:2360-6. [PubMed]
- Rogenhofer S, Kahl P, Mertens C, et al. Global histone H3 lysine 27 (H3K27) methylation levels and their prognostic relevance in renal cell carcinoma. BJU Int 2012;109:459-65. [PubMed]
- Xu B, Abourbih S, Sircar K, et al. Enhancer of zeste homolog 2 expression is associated with metastasis and adverse clinical outcome in clear cell renal cell carcinoma: a comparative study and review of the literature. Arch Pathol Lab Med 2013;137:1326-36. [PubMed]
- Wagener N, Macher-Goeppinger S, Pritsch M, et al. Enhancer of zeste homolog 2 (EZH2) expression is an independent prognostic factor in renal cell carcinoma. BMC Cancer 2010;10:524. [PubMed]
- Hausser J, Zavolan M. Identification and consequences of miRNA-target interactions--beyond repression of gene expression. Nat Rev Genet 2014;15:599-612. [PubMed]
- Lim LP, Lau NC, Garrett-Engele P, et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005;433:769-73. [PubMed]
- Lewis BP, Shih IH, Jones-Rhoades MW, et al. Prediction of mammalian microRNA targets. Cell 2003;115:787-98. [PubMed]
- Iorio MV, Croce CM. MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol Med 2012;4:143-59. [PubMed]
- Chow TF, Mankaruos M, Scorilas A, et al. The miR-17-92 cluster is over expressed in and has an oncogenic effect on renal cell carcinoma. J Urol 2010;183:743-51. [PubMed]
- Valera VA, Walter BA, Linehan WM, et al. Regulatory Effects of microRNA-92 (miR-92) on VHL Gene Expression and the Hypoxic Activation of miR-210 in Clear Cell Renal Cell Carcinoma. J Cancer 2011;2:515-26. [PubMed]
- Lichner Z, Mejia-Guerrero S, Ignacak M, et al. Pleiotropic action of renal cell carcinoma-dysregulated miRNAs on hypoxia-related signaling pathways. Am J Pathol 2012;180:1675-87. [PubMed]
- McCormick RI, Blick C, Ragoussis J, et al. miR-210 is a target of hypoxia-inducible factors 1 and 2 in renal cancer, regulates ISCU and correlates with good prognosis. Br J Cancer 2013;108:1133-42. [PubMed]
- Nakada C, Tsukamoto Y, Matsuura K, et al. Overexpression of miR-210, a downstream target of HIF1α, causes centrosome amplification in renal carcinoma cells. J Pathol 2011;224:280-8. [PubMed]
- Hirata H, Hinoda Y, Ueno K, et al. MicroRNA-1826 directly targets beta-catenin (CTNNB1) and MEK1 (MAP2K1) in VHL-inactivated renal cancer. Carcinogenesis 2012;33:501-8. [PubMed]
- Hell MP, Thoma CR, Fankhauser N, et al. miR-28-5p promotes chromosomal instability in VHL-associated cancers by inhibiting Mad2 translation. Cancer Res 2014;74:2432-43. [PubMed]
- Pencheva N, Tavazoie SF. Control of metastatic progression by microRNA regulatory networks. Nat Cell Biol 2013;15:546-54. [PubMed]
- Bera A, Das F, Ghosh-Choudhury N, et al. microRNA-21-induced dissociation of PDCD4 from rictor contributes to Akt-IKKβ-mTORC1 axis to regulate renal cancer cell invasion. Exp Cell Res 2014;328:99-117. [PubMed]
- Xia H, Hui KM. MicroRNAs involved in regulating epithelial-mesenchymal transition and cancer stem cells as molecular targets for cancer therapeutics. Cancer Gene Ther 2012;19:723-30. [PubMed]
- Yoshino H, Enokida H, Itesako T, et al. Epithelial-mesenchymal transition-related microRNA-200s regulate molecular targets and pathways in renal cell carcinoma. J Hum Genet 2013;58:508-16. [PubMed]
- Korpal M, Lee ES, Hu G, et al. The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 2008;283:14910-4. [PubMed]
- Chen X, Wang X, Ruan A, et al. miR-141 is a key regulator of renal cell carcinoma proliferation and metastasis by controlling EphA2 expression. Clin Cancer Res 2014;20:2617-30. [PubMed]
- Huang J, Yao X, Zhang J, et al. Hypoxia-induced downregulation of miR-30c promotes epithelial-mesenchymal transition in human renal cell carcinoma. Cancer Sci 2013;104:1609-17. [PubMed]
- Yamasaki T, Seki N, Yamada Y, et al. Tumor suppressive microRNA-138 contributes to cell migration and invasion through its targeting of vimentin in renal cell carcinoma. Int J Oncol 2012;41:805-17. [PubMed]
- Yamasaki T, Seki N, Yoshino H, et al. MicroRNA-218 inhibits cell migration and invasion in renal cell carcinoma through targeting caveolin-2 involved in focal adhesion pathway. J Urol 2013;190:1059-68. [PubMed]
- Zen K, Zhang CY. Circulating microRNAs: a novel class of biomarkers to diagnose and monitor human cancers. Med Res Rev 2012;32:326-48. [PubMed]
- Mitchell PS, Parkin RK, Kroh EM, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A 2008;105:10513-8. [PubMed]
- Iwamoto H, Kanda Y, Sejima T, et al. Serum miR-210 as a potential biomarker of early clear cell renal cell carcinoma. Int J Oncol 2014;44:53-8. [PubMed]
- Redova M, Poprach A, Nekvindova J, et al. Circulating miR-378 and miR-451 in serum are potential biomarkers for renal cell carcinoma. J Transl Med 2012;10:55. [PubMed]
- Wang C, Hu J, Lu M, et al. A panel of five serum miRNAs as a potential diagnostic tool for early-stage renal cell carcinoma. Sci Rep 2015;5:7610. [PubMed]
- Mlcochova H, Hezova R, Stanik M, et al. Urine microRNAs as potential noninvasive biomarkers in urologic cancers. Urol Oncol 2014;32:41.e1-9.
- Yun SJ, Jeong P, Kim WT, et al. Cell-free microRNAs in urine as diagnostic and prognostic biomarkers of bladder cancer. Int J Oncol 2012;41:1871-8. [PubMed]
- von Brandenstein M, Pandarakalam JJ, Kroon L, et al. MicroRNA 15a, inversely correlated to PKCα, is a potential marker to differentiate between benign and malignant renal tumors in biopsy and urine samples. Am J Pathol 2012;180:1787-97. [PubMed]
- Zaman MS, Shahryari V, Deng G, et al. Up-regulation of microRNA-21 correlates with lower kidney cancer survival. PLoS One 2012;7:e31060. [PubMed]
- Fritz HK, Lindgren D, Ljungberg B, et al. The miR(21/10b) ratio as a prognostic marker in clear cell renal cell carcinoma. Eur J Cancer 2014;50:1758-65. [PubMed]
- Zhao X, Zhao Z, Xu W, et al. Down-regulation of miR-497 is associated with poor prognosis in renal cancer. Int J Clin Exp Pathol 2015;8:758-64. [PubMed]
- Ishihara T, Seki N, Inoguchi S, et al. Expression of the tumor suppressive miRNA-23b/27b cluster is a good prognostic marker in clear cell renal cell carcinoma. J Urol 2014;192:1822-30. [PubMed]
- Samaan S, Khella HW, Girgis A, et al. miR-210 is a prognostic marker in clear cell renal cell carcinoma. J Mol Diagn 2015;17:136-44. [PubMed]
- Mangolini A, Bonon A, Volinia S, et al. Differential expression of microRNA501-5p affects the aggressiveness of clear cell renal carcinoma. FEBS Open Bio 2014;4:952-65. [PubMed]
- Khella HW, Scorilas A, Mozes R, et al. Low expression of miR-126 is a prognostic marker for metastatic clear cell renal cell carcinoma. Am J Pathol 2015;185:693-703. [PubMed]
- Wotschofsky Z, Busch J, Jung M, et al. Diagnostic and prognostic potential of differentially expressed miRNAs between metastatic and non-metastatic renal cell carcinoma at the time of nephrectomy. Clin Chim Acta 2013;416:5-10. [PubMed]
- Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet 2009;10:155-9. [PubMed]
- Ravasi T, Suzuki H, Pang KC, et al. Experimental validation of the regulated expression of large numbers of non-coding RNAs from the mouse genome. Genome Res 2006;16:11-9. [PubMed]
- Kornienko AE, Guenzl PM, Barlow DP, et al. Gene regulation by the act of long non-coding RNA transcription. BMC Biol 2013;11:59. [PubMed]
- Tripathi V, Ellis JD, Shen Z, et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell 2010;39:925-38. [PubMed]
- Yang L, Lin C, Liu W, et al. ncRNA- and Pc2 methylation-dependent gene relocation between nuclear structures mediates gene activation programs. Cell 2011;147:773-88. [PubMed]
- Wu XS, Wang XA, Wu WG, et al. MALAT1 promotes the proliferation and metastasis of gallbladder cancer cells by activating the ERK/MAPK pathway. Cancer Biol Ther 2014;15:806-14. [PubMed]
- Zhang HM, Yang FQ, Chen SJ, et al. Upregulation of long non-coding RNA MALAT1 correlates with tumor progression and poor prognosis in clear cell renal cell carcinoma. Tumour Biol 2015;36:2947-55. [PubMed]
- Yang F, Zhang L, Huo XS, et al. Long noncoding RNA high expression in hepatocellular carcinoma facilitates tumor growth through enhancer of zeste homolog 2 in humans. Hepatology 2011;54:1679-89. [PubMed]
- Kim K, Jutooru I, Chadalapaka G, et al. HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene 2013;32:1616-25. [PubMed]
- Lv XB, Lian GY, Wang HR, et al. Long noncoding RNA HOTAIR is a prognostic marker for esophageal squamous cell carcinoma progression and survival. PLoS One 2013;8:e63516. [PubMed]
- Wu Y, Liu J, Zheng Y, et al. Suppressed expression of long non-coding RNA HOTAIR inhibits proliferation and tumourigenicity of renal carcinoma cells. Tumour Biol 2014;35:11887-94. [PubMed]
- Chiyomaru T, Fukuhara S, Saini S, et al. Long non-coding RNA HOTAIR is targeted and regulated by miR-141 in human cancer cells. J Biol Chem 2014;289:12550-65. [PubMed]
- Zhang HM, Yang FQ, Yan Y, et al. High expression of long non-coding RNA SPRY4-IT1 predicts poor prognosis of clear cell renal cell carcinoma. Int J Clin Exp Pathol 2014;7:5801-9. [PubMed]
- Qiao HP, Gao WS, Huo JX, et al. Long non-coding RNA GAS5 functions as a tumor suppressor in renal cell carcinoma. Asian Pac J Cancer Prev 2013;14:1077-82. [PubMed]
- Murakami Y. Involvement of a cell adhesion molecule, TSLC1/IGSF4, in human oncogenesis. Cancer Sci 2005;96:543-52. [PubMed]
- Yao J, Chen Y, Wang Y, et al. Decreased expression of a novel lncRNA CADM1-AS1 is associated with poor prognosis in patients with clear cell renal cell carcinomas. Int J Clin Exp Pathol 2014;7:2758-67. [PubMed]
- Malouf GG, Zhang J, Yuan Y, et al. Characterization of long non-coding RNA transcriptome in clear-cell renal cell carcinoma by next-generation deep sequencing. Mol Oncol 2015;9:32-43. [PubMed]
- Mattei J, da Silva RD, Sehrt D, et al. Targeted therapy in metastatic renal carcinoma. Cancer Lett 2014;343:156-60. [PubMed]


